From February 6th to February 10th, France-BioImaging organised a group meeting on the project “FBI.data” in Bordeaux. For a week, participants focused on the architecture and the implementation of image data management tools. A user-friendly response to the challenges of never-ending data production.
New imaging technologies are very greedy in terms of image processing and data management. Beside the image itself, biological imaging generates a huge amount of metadata. The FBI.data project, one of the key missions of France-BioImaging, addresses the questions related to the computational analysis and handling of image data.
Speeding up the implementation of tools across the infrastructure
Although the distributed FBI.data team meets once per week, the FBI.data Sprint aims at only focusing on data management scenarii and accelerating the project. Two essential aspects have been discussed:
- First of all, the data management system architecture must be simple for them to be implemented across France-BioImaging nodes. It also has to be compatible with long-term data storage and of course, to be user-friendly – we want to keep it easy for our users!
- The second point is all about anticipating as many data management cases as possible. Running through all the needs of bioimaging experts and users, the team lists the specific features of each case and considers the perfect solutions for all of them.
Working for the FAIRisation of data
The FAIRisation of data for Open Science is an initiative fully endorsed by France-BioImaging. Meaning that data are Findable, Accessible, Interoperable and Reusable, the benefits for the bioimaging community are numerous. It improves transparency and reproducibility, enhances quality of results, accelerates scientific progress and method development and finally boosts collaboration within the scientific community.
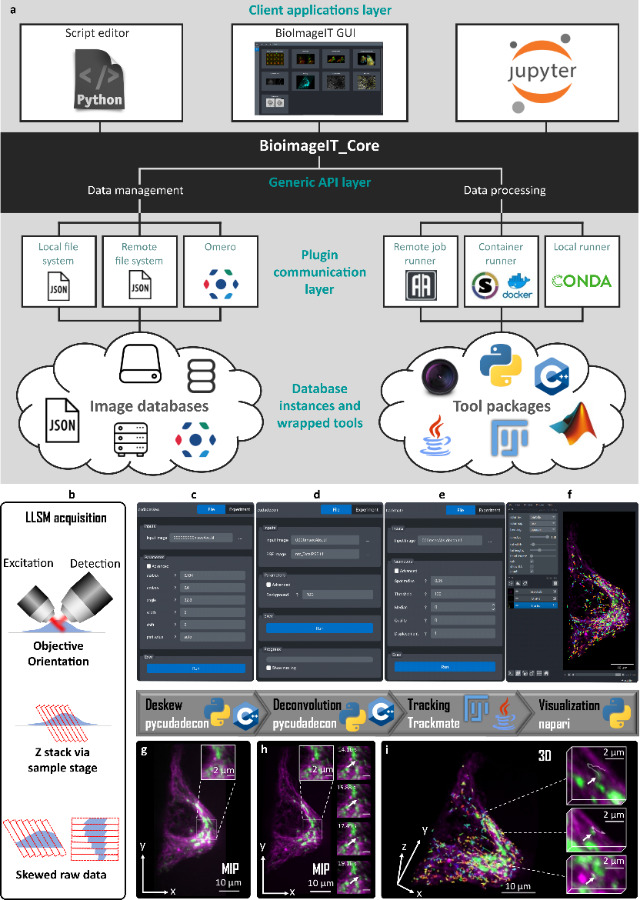
OMERO, developed by the University of Dundee & Open Microscopy Environment teams, on which the FBI.data group is working, is one of the software making user data FAIR. Being a microscopy image data management decentralised platform, it helps organise, access and archive data. Besides, it combines image and metadata storage, a viewer and data analysis resources. Furthermore, OMERO is linked to the most valuable tools for bioimaging experts (ImageJ, Napari, QPath, etc.). And users can access their data from anywhere and keep them safe.
But much more has to be set up to have functional solutions: to ease user authentication and management, manage big data transfer, and have an adequate metadata scheme. Accompanying users is one of the mission of the FBI.data team, and the FBI.data Sprint is also the occasion to join efforts from the training working group led by the training mission officer Fabrice Cordelières (Bordeaux Imaging Center) to produce adequate training material on data management.
Sharing efforts and helping the community
The FBI.data working group is composed of:
- Perrine Paul-Gilloteaux, research engineer at MicroPICell core facility and FBI.data mission officer
- Emmanuel Faure, researcher at LIRMM (Laboratoire d’informatique, de robotique et de microélectronique de Montpellier) and FBI.data mission officer
- Guillaume Gay, research engineer at LIRMM, working full time for the FBI.data project
- Marc Mongy, research engineer at MicroPIcell, working full time for the FBI.data project
- Guillaume Maucort, research engineer in image analysis at Bordeaux Imaging Center (BIC)
- Jean-François Guillaume, research Engineer on the BIRD facility, Nantes and Pays de La Loire mesocenter (shared IT system engineer between FBI and Institut Français de Bioinformatique)
- Thierry Pecot, research engineer in image analysis at Rennes (Bretagne Loire Node)
- Further recruitments are on-going and will be reinforcing the team
By joining their skills and experience, they are working together on setting up tools and good practices for the management and FAIRisation of data inside France-BioImaging nodes but also for the entire bioimaging community. With this in mind, the project has collaboration with, among others, the Institut Français de Bioinformatique (IFB) and the Centre National de Ressources Biologiques Marines (EMBRC), and other infrastructure through the MUDIS4LS Equipex+ project. Moreover, the FBI.data project has an open GitLab, providing image data management codes in open source, and a blog with tutorials, recommendations and so much more!
Check the France-BioImaging OMERO web portal: https://omero-fbi.fr/
And its gitlab FBI data · GitLab (in2p3.fr)
Learn more about our missions and working packages: https://france-bioimaging.org/about/work-packages/