Cryogenic electron tomography (cryo-ET) visualizes the 3D spatial distribution of macromolecules at nanometer resolution inside native cells. However, automated identification of macromolecules inside cellular tomograms is challenged by noise and reconstruction artifacts, as well as the presence of many molecular species in the crowded volumes.
To overcome these obstacles, an international team of scientists from France, Spain and Germany, under the leadership of Charles Kervrann, from France BioImaging BioImage Informatics Node, developed a deep learning-based framework to quickly identify multiple classes of macromolecules in cryo-ET volumes. This DeepFinder program, now published in Nature Methods, builds upon convolutional neural networks that have already proven highly valuable in the microscopy field.
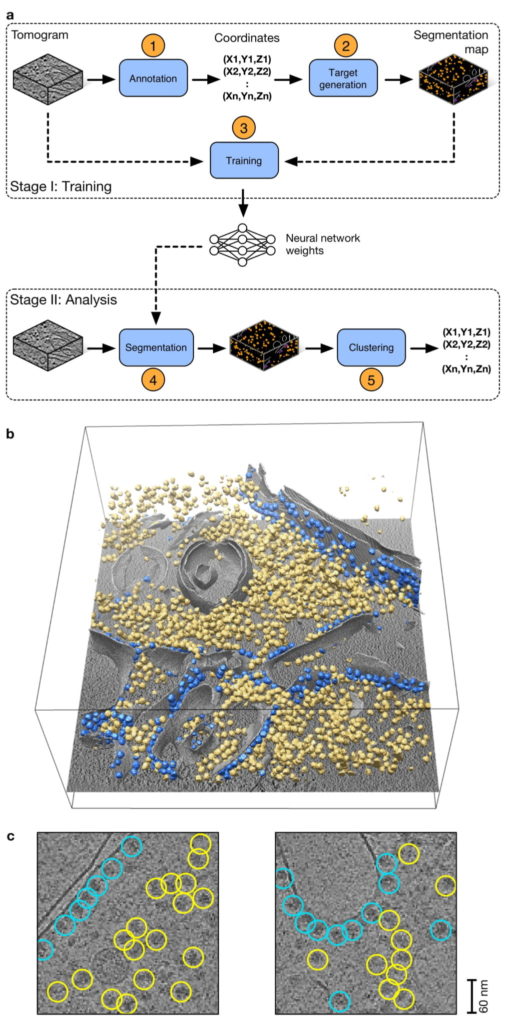
Overview of DeepFinder (from Moebel, E., et al., Nat Methods 18, 1386–1394 (2021).
a) The DeepFinder workflow consists of a training stage (stage I) and an analysis (or inference) stage (stage II). These two stages correspond to five steps (represented by blue boxes) to locate macromolecular complexes within crowded cells.
b) Ribosome localization with DeepFinder in a cryo-electron tomogram of a C. reinhardtii cell.
Tomographic slice with superimposed segmented cell membrane (gray) and ribosomes classified with respect to their binding states: membrane-bound (blue) and cytosolic (yellow).
c) Tomographic slices showing coordinates
of detected ribosomes (colors correspond to b). The positions and classes were determined by analyzing the segmentation map shown in b. This
analysis used 48 tomograms for training, one for validation and eight for testing. Scale bar, 60 nm.
Once trained, the inference stage of DeepFinder is faster than template matching and performs better than other competitive deep learning methods at identifying macromolecules of various sizes in both synthetic and experimental datasets. On cellular cryo-ET data, DeepFinder localized membrane-bound and cytosolic ribosomes (roughly 3.2 MDa), ribulose 1,5-bisphosphate carboxylase–oxygenase (roughly 560 kDa soluble complex) and photosystem II (roughly 550 kDa membrane complex) with an accuracy comparable to expert-supervised ground truth annotations. DeepFinder is therefore a promising algorithm for the semiautomated analysis of a wide range of molecular targets in cellular tomograms. It also serves as a prime example illustrating the importance of developing efficient, customized AI tools to accelerate knowledge generation in the biomedical life sciences.
DeepFinder has been implemented as a free, open-source program with an accessible graphical user interface.
The team is currently working on adapting it to fluorescence microscopy.
Moebel, E., Martinez-Sanchez, A., Lamm, L. et al. Deep learning improves macromolecule identification in 3D cellular cryo-electron tomograms. Nat Methods 18, 1386–1394 (2021). https://doi.org/10.1038/s41592-021-01275-4