A new version of TrackMate is available now, with major changes that improve its versatility. TrackMate now integrates state-of-the-art segmentation algorithms from machine-learning and deep-learning such as StarDist, Ilastik and Weka.
TrackMate[1] is a Fiji plugin that address cell or organelle tracking in Life-Science microscopy images. Its main goals are to be user-friendly, interoperable and to serve as a platform to accelerate the development of novel tracking algorithms and analysis pipelines.
With this new version we rewrote almost entirely TrackMate so that it can integrate state-of-the-art segmentation algorithms and benefit from their output. For instance, TrackMate can now store, display, save, load and exploit object contours in 2D.
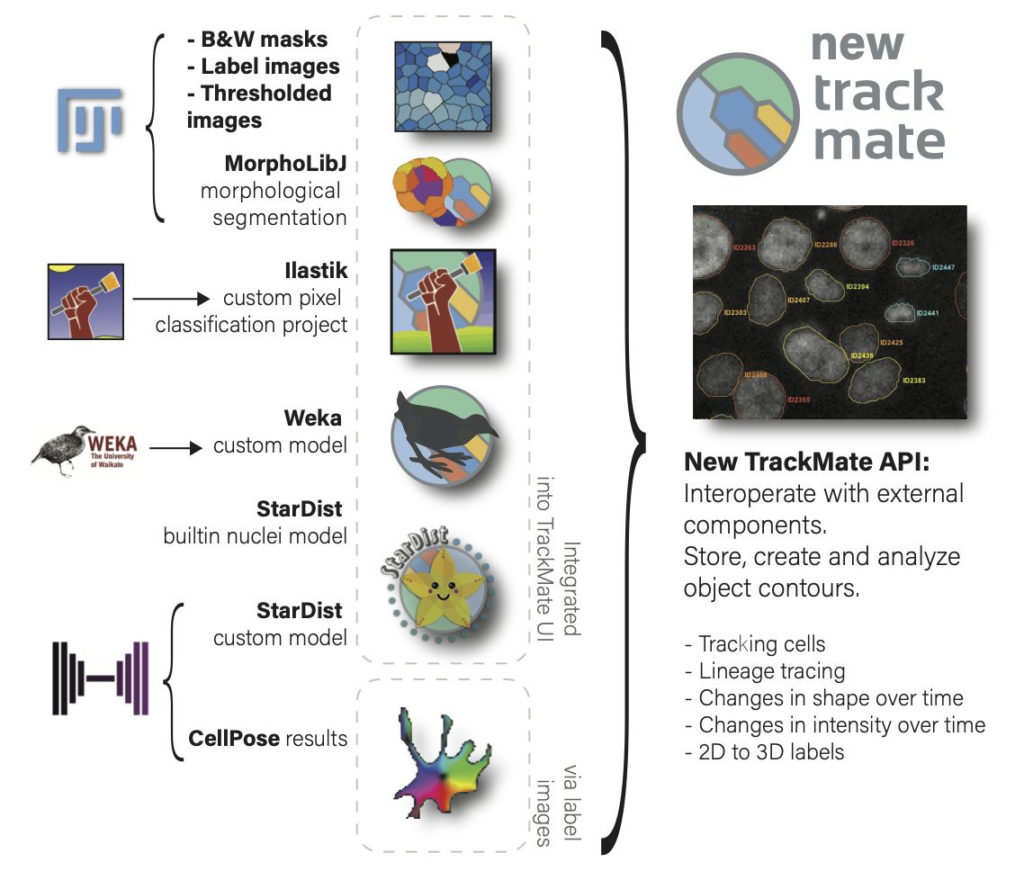
We also made a new application programming interface that will facilitate and accelerate reusing TrackMate in other analysis pipelines and allow 3rd party contributors to add new segmentation algorithms in TrackMate in an easy way. We used this API ourselves to add 7 new segmentation algorithms to TrackMate:
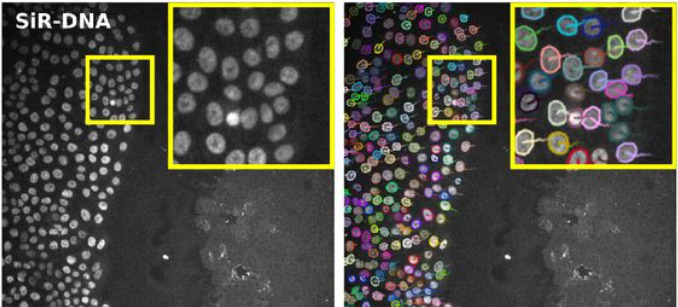
For instance, the StarDist[2] algorithm is integrated as two different detectors. The first one uses the built-in deep-learning model that can segment cell nuclei in fluorescence image in a wide range of situation. The robustness of the StarDist algorithm in turn positively impacts the robustness of tracking and allows for better detection of cell divisions with TrackMate tracking algorithms. This will facilitate cell migration studies.
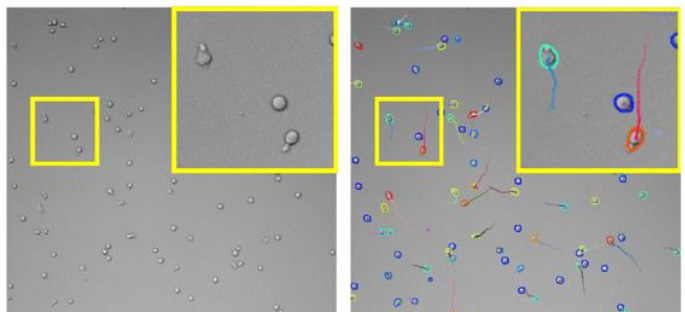
The TrackMate StarDist integration also allows for specifying and using a custom deep-learning model. For instance, we trained a specific model to detect T-cells imaged in bright-field microscopy and track them over time. Before the emergence of such detection algorithms, the tracking of label-free cells was difficult.

We also integrated the ilastik[3] segmentation software. A TrackMate user can input an ilastik classifier to detect objects then track them. We used them to study the bacterial growth of Neisseria meningitidis clones. The output of this analysis pipeline offers the lineage of each single cell along with its morphology and how it evolves across cell divisions.
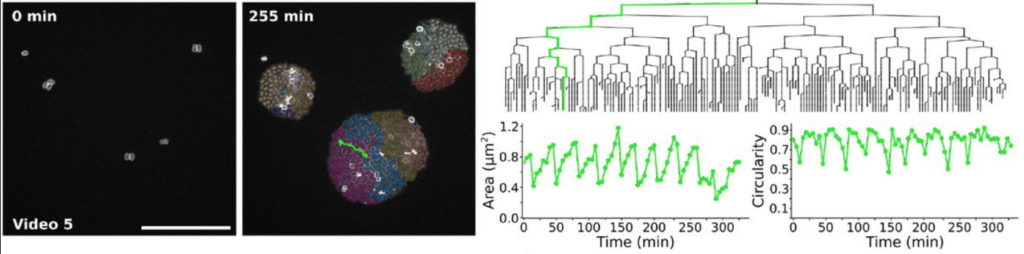
The new capabilities of TrackMate can be used to address applications beyond tracking. For instance, it is now possible to use TrackMate to perform the segmentation of 3D objects using a slice-by-slice approach. This approach consists in segmenting objects in each 2D section of a 3D stack, then merging the segmentation results along Z in a subsequent step. This can be done in TrackMate, using the tracking step for merging. We implemented a novel tracking algorithm to foster this application, the overlap tracker. We could use this approach combining the cellpose[4] algorithm in 2D to segment 3D images of Arabidopsis thaliana floral meristem.
There are several other algorithms that are now offered to the TrackMate user, within a user-friendly software meant to interoperate with the key software of bioimage analysis. More importantly, TrackMate is an open-source academic software, and its new API will foster the development of new analysis pipeline with TrackMate and the integration of new algorithms by other developers, increasing the breadth of applications it can address for Life-Science researchers.
This new version of TrackMate is the product of a collaboration between the IAH facility (Institut Pasteur), part of the FBI Bioimage Informatics Node , the Jacquemet lab (Turku Bioscience Centre) , and the Dumenil lab (Institut Pasteur) .
Bringing TrackMate in the era of machine-learning and deep-learningDmitry Ershov, Minh-Son Phan, Joanna W. Pylvänäinen, Stéphane U. Rigaud, Laure Le Blanc, Arthur Charles-Orszag, James R. W. Conway, Romain F. Laine, Nathan H. Roy, Daria Bonazzi, Guillaume Duménil, Guillaume Jacquemet, Jean-Yves Tinevez bioRxiv 2021.09.03.458852; doi: https://doi.org/10.1101/2021.09.03.458852
Contact: Jean-Yves Tinevez
[1] https://imagej.net/plugins/trackmate/ [2] Alejandro F. Frangi, Julia A. Schnabel, Christos Davatzikos, Carlos Alberola-López, and Gabor Fichtinger Uwe Schmidt, Martin Weigert, Coleman Broaddus, and Gene Myers. Cell detection with star-convex polygons. In Alejandro F. Frangi, Julia A. Schnabel, Christos Davatzikos, Carlos Alberola-López, and Gabor Fichtinger, editors, Medical Image Computing and Computer Assisted Intervention – MICCAI 2018, pages 265–273, Cham, 2018. Springer International Publishing. doi:10.1007/978-3-030-00934-2_30. [3] Stuart Berg, Dominik Kutra, Thorben Kroeger, Christoph N Straehle, Bernhard X Kausler, Carsten Haubold, Martin Schiegg, Janez Ales, Thorsten Beier, Markus Rudy, Kemal Eren, Jaime I Cervantes, Buote Xu, Fynn Beuttenmueller, Adrian Wolny, Chong Zhang, Ullrich Koethe, Fred A Hamprecht, and Anna Kreshuk. ilastik: interactive machine learning for (bio)image analysis. Nature Methods, 16(12):1226–1232, 2019. ISSN 1548-7105. doi:10.1038/s41592-019-0582-9.
[4] Carsen Stringer, Tim Wang, Michalis Michaelos, and Marius Pachitariu. Cellpose: a generalist algorithm for cellular segmentation. Nature Methods, 18(1):100–106, jan 2021. doi:10.1038/s41592-020-01018-x.

