The PICsL-FBI microscopy core facility is located on two sites: Centre d’Immunologie de Marseille Luminy (CIML) and Institut de Biologie du Développement de Marseille (IBDM).The PICsL-FBI facility of the CIML called ImagImm (Imaging Immunity) via its microscopy resources – from the molecule to whole organisms – is dedicated to help its users deciphering cellular mechanisms in the fields of immunology.
Major research implications of the ImagImm facility:
- Technology transfer: spot variation Fluorescence Correlation Spectroscopy (svFCS)
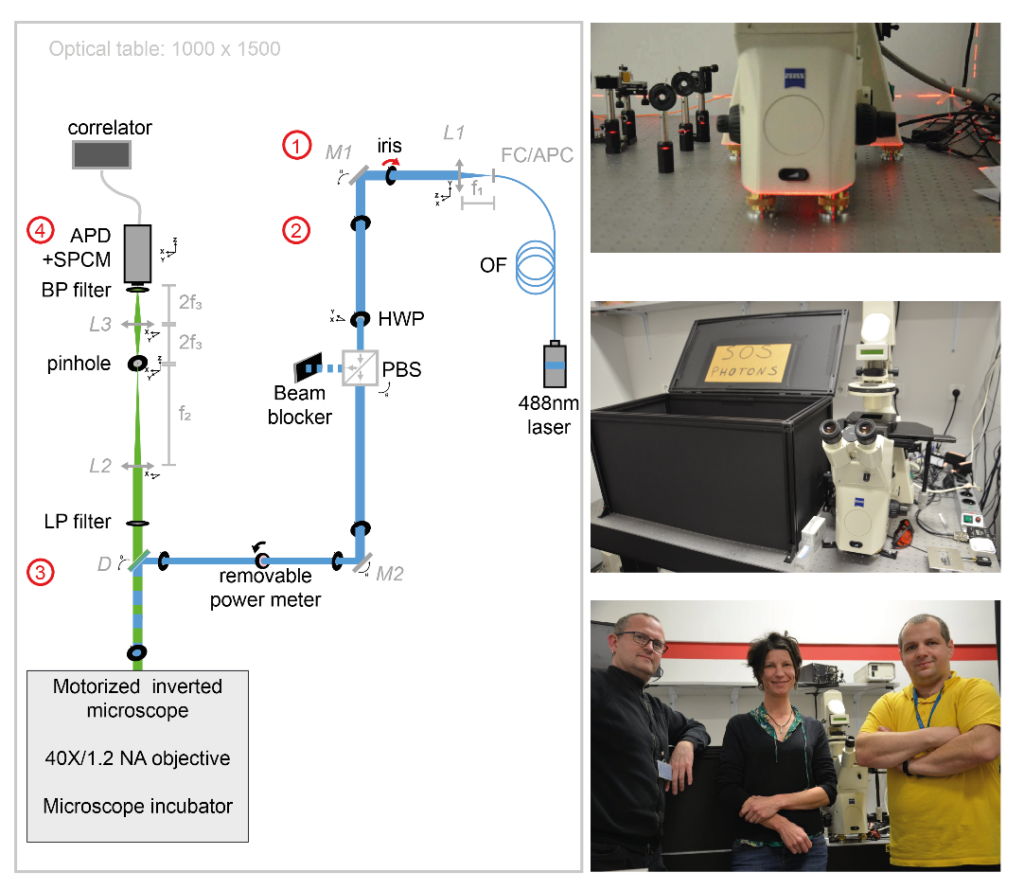
In collaboration with Tomasz Trombik (Faculty of Biotechnology, University of Wroclaw – Wroclaw, Poland), Sophie Brustlein (Institut de Convergences Centuri, AMU,CNRS – Marseille, France) and Nicolas Bertaux (Institut Fresnel, AMU, Centrale Marseille, CNRS – Marseille, France), Sébastien Mailfert and Didier Marguet published the procedure for implementing spot variation Fluorescence Correlation Spectroscopy (svFCS) measurements using a classical fluorescence microscope that has been customized1. This publication is following the technology transfer made in 2018: the svFCS developed by Didier Marguet’s lab was duplicated by Sébastien Mailfert and Sophie Brustlein and built from scratch in 7 days on site, in Poland.
Dynamic biological processes in living cells, including those associated with plasma membrane organization, occur on various spatial and temporal scales, ranging from nanometers to micrometers and microseconds to minutes, respectively. Such a broad range of biological processes challenges conventional microscopy approaches. The published protocol includes a specific performance check of the svFCS setup and the guidelines for molecular diffusion measurements by svFCS on the plasma membrane of living cells under physiological conditions. Additionally, a procedure for disrupting plasma membrane raft nanodomains by cholesterol oxidase treatment is provided and how these changes in the lateral organization of the plasma membrane might be revealed by svFCS analysis. This fluorescence-based method can provide unprecedented details on the lateral organization of the plasma membrane with the appropriate spatial and temporal resolution.
- SAPHIR : a Shiny application to analyze tissue section images
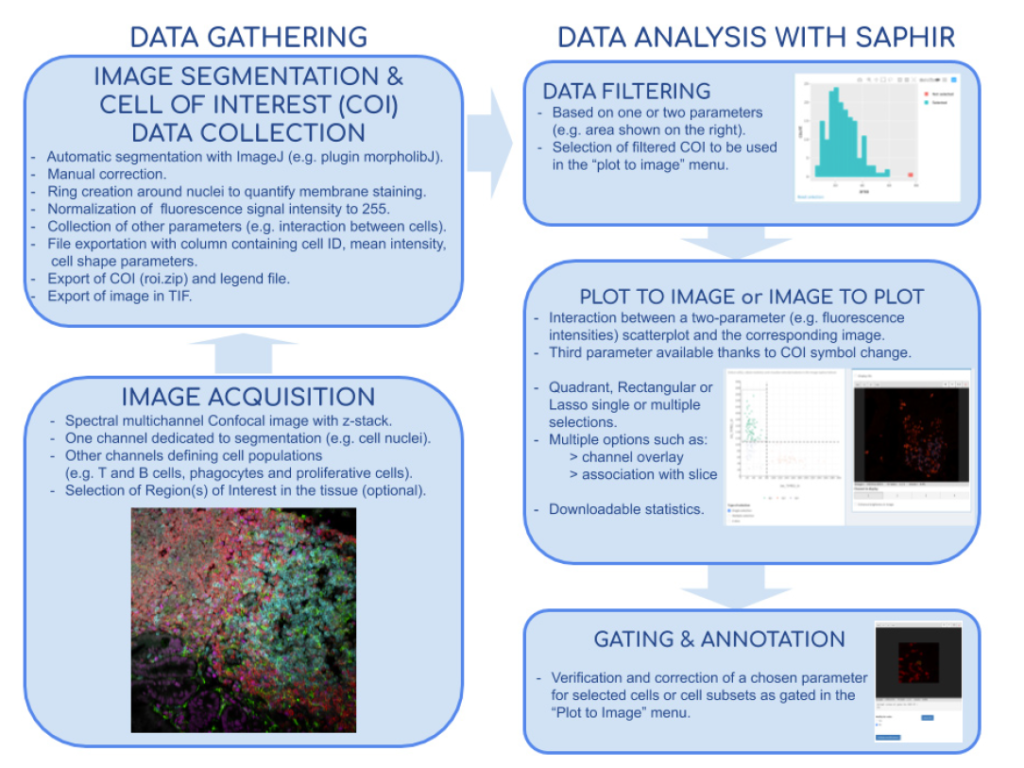
In collaboration with Hugues Lelouard (CIML, Inserm, CNRS, AMU) and Elodie Germani, Mathieu Fallet published a powerful method for both basic and medical research to study cell populations in tissues using immunofluorescence. Image acquisitions performed by confocal microscopy notably allow excellent lateral resolution and more than 10 parameter measurement when using spectral or multiplexed imaging. Analysis of such complex images can be very challenging and easily lead to bias and misinterpretation. They developed the Shiny Analytical Plot of Histological Images Results (SAPHIR), an R shiny application for histo-cytometry using scatterplot representation of data extracted by segmentation. It offers many features, such as filtering of spurious data points, selection of cell subsets on scatterplot, visualization of scatterplot selections back into the image, statistics of selected data and data annotation. This application allows to quickly characterize labeled cells, from their phenotype to their number and location in the tissue, as well as their interaction with other cells.
- Wound healing in C. elegans
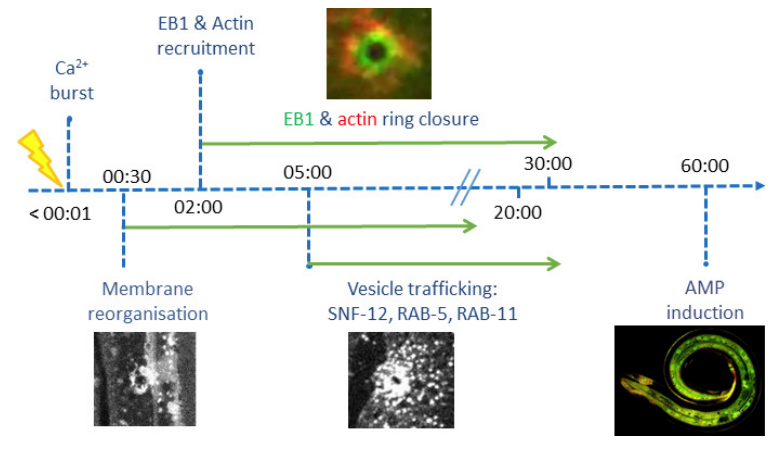
In collaboration with Nathalie Pujol and Jonathan Ewbank (CIML, Inserm, CNRS, AMU), Mathieu Fallet and Sébastien Mailfert participated in the project on the immune response by showing that wounding provokes a reorganization of plasma membrane subdomains3. The skin protects animals from infection and physical damage. In Caenorhabditis elegans, wounding the epidermis triggers an immune reaction and a repair response, but it is not clear how these are coordinated. Previous work implicated the microtubule cytoskeleton in the maintenance of epidermal integrity (Chuang et al., 2016). Taffoni et al. show the reorganization of the plasma membrane subdomains by a simple wounding system. This is followed by recruitment of the microtubule plus end-binding protein EB1/EBP-2 around the wound and actin ring formation, dependent on ARP2/3 branched actin polymerization. They show that microtubule dynamics are required for the recruitment and closure of the actin ring, and for the trafficking of the key signaling protein SLC6/SNF-12 toward the injury site. Without SNF-12 recruitment, there is an abrogation of the immune response. These results suggest that microtubule dynamics coordinate the cytoskeletal changes required for wound repair and the concomitant activation of innate immunity.
References:
- Mailfert, S., Wojtowicz, K., Brustlein, S., Blaszczak, E., Bertaux, N., Łukaszewicz, M., Marguet, D., Trombik, T. Spot Variation Fluorescence Correlation Spectroscopy for Analysis of Molecular Diffusion at the Plasma Membrane of Living Cells, JoVE, 165, 1-19 (2020).
- Germani, E., Lelouard, H., Fallet, M. SAPHIR: a Shiny application to analyze tissue section images, F1000Research, Faculty of 1000, 9, 1276-1285 (2020).
- Taffoni, C., Omi, S., Huber, C., Mailfert, S., Fallet, M., Rupprecht, J-F,. Ewbank, J., Pujol., N. Microtubule plus-end dynamics link wound repair to the innate immune response, eLIFE, 9, e45047 (2020)