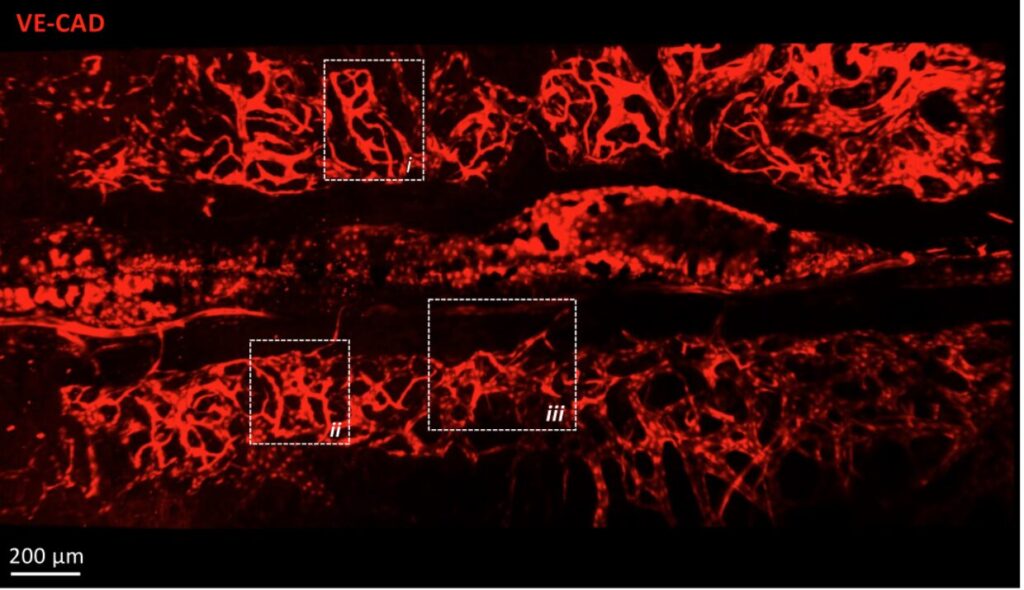
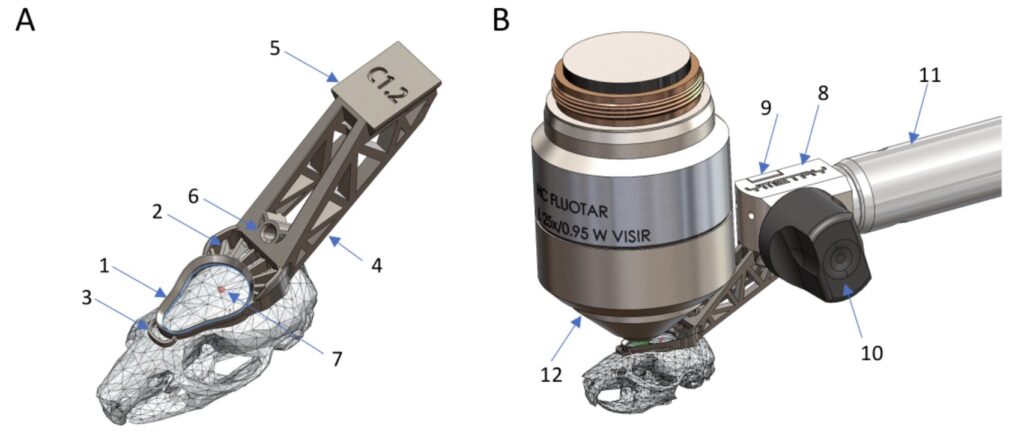
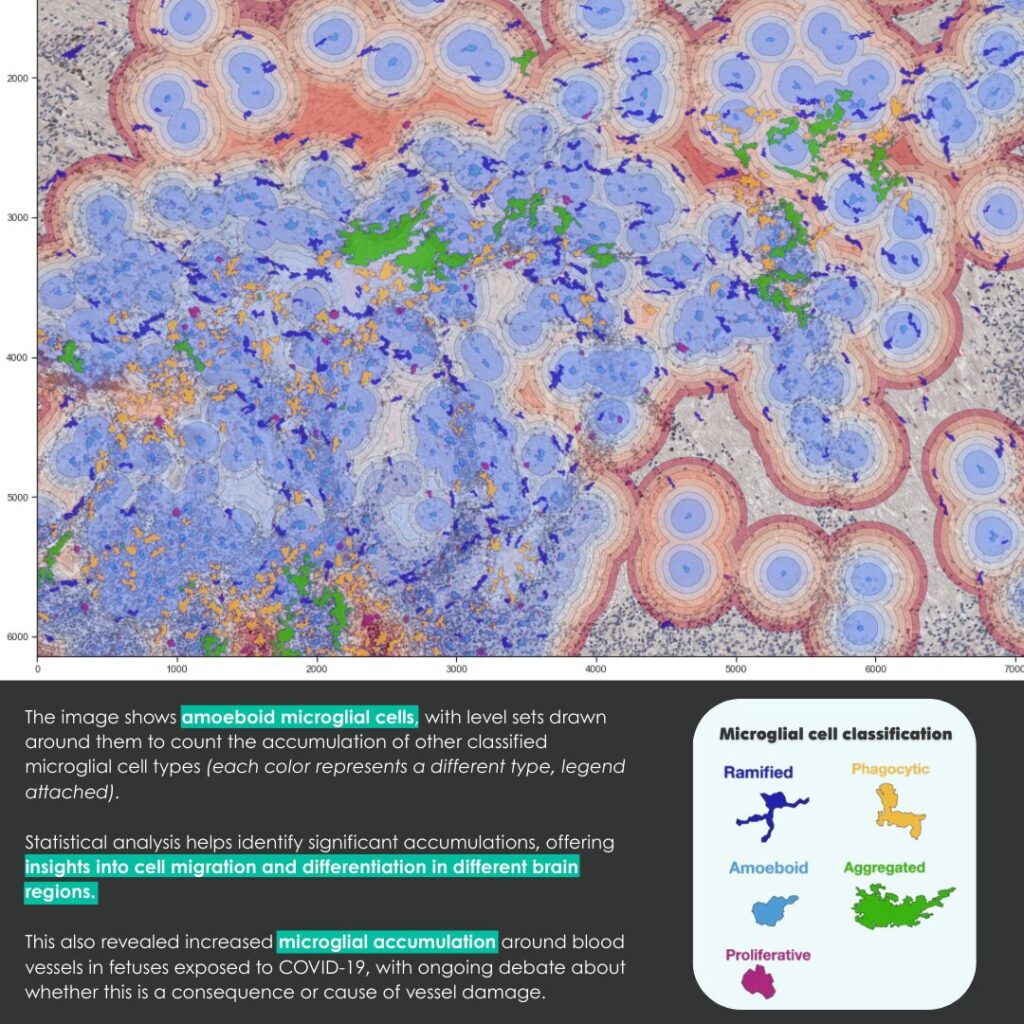
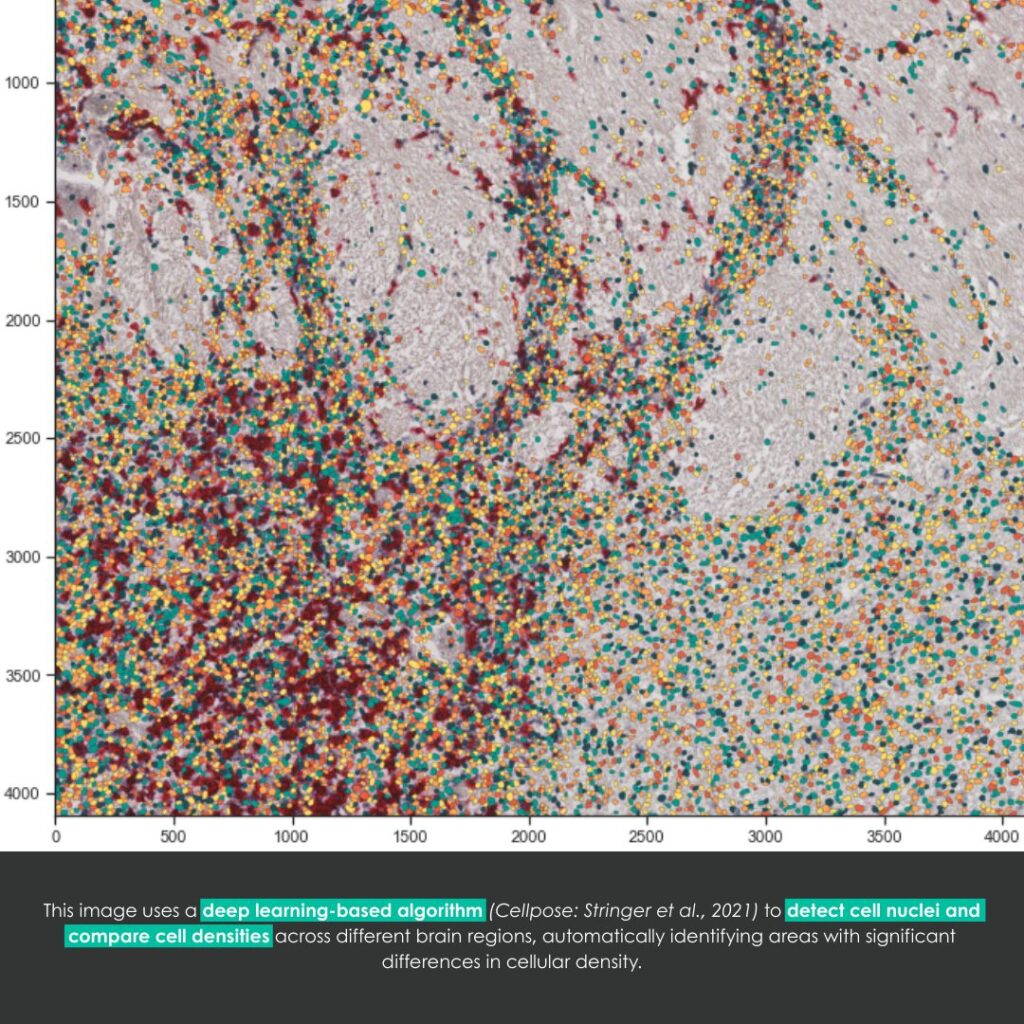
Fluorescence microscopy allows researchers to explore the living world at the cellular and subcellular scales with remarkable precision. However, as time passes, microscopes inevitably degrade: detectors become noisy, optical systems lose alignment, and image quality declines. This aging process can hinder long-term biological studies and quantitative analysis.
To address this challenge, a team of Engineers from IBDM and LIS (France-BioImaging Marseille node) developed μPIX, a new deep-learning algorithm based on generative artificial intelligence.
A smarter way to restore microscopy images
μPIX uses a specific type of AI called a Pix2Pix conditional Generative Adversarial Network (cGAN): this algorithm learns how to transform low-quality or noisy images into clean and high-quality ones, based on examples.
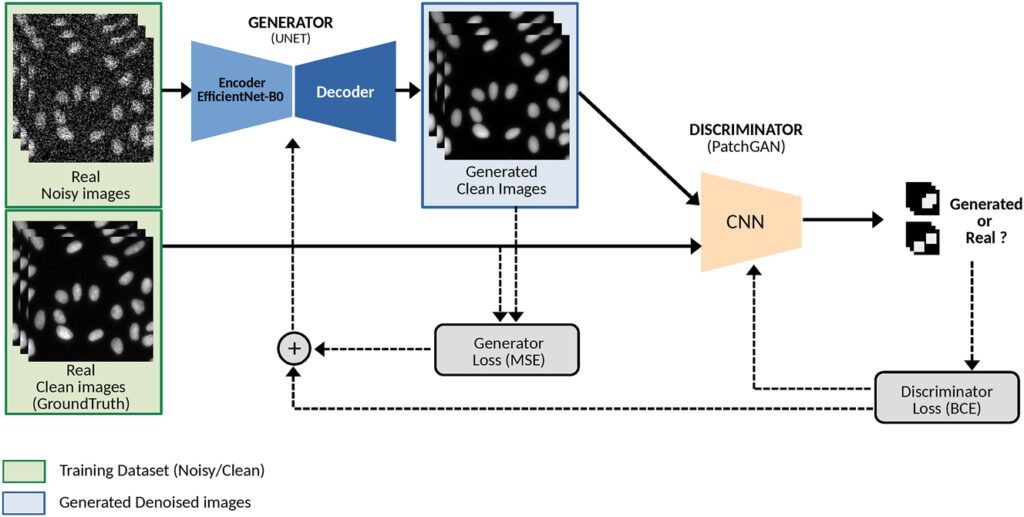
Figure: µPIX architecture is based on a Pix2Pix generative network. µPIX consists of two subnetworks: a generator, based on a UNet architecture with an EfficientNet-b0 backbone, and a discriminator (PatchGAN). During supervised training, a noisy image is input to the generator, which generate an image. This output is compared to the real clean image using a pixel-wise loss function (MSE). Pairs of real and generated images are then passed to the discriminator, which classifies them as real or fake using a binary cross-entropy loss (BCE). Both subnetworks are progressively refined through adversarial loss during training. In the inference phase, only the trained generator is used to generate clean images. (Bon, Gabriel, Sapède, Daniel, Matthews, Cédric and Daian, Fabrice. “μPIX: leveraging generative AI for enhanced, personalized and sustainable microscopy” Methods in Microscopy, 2025. https://doi.org/10.1515/mim-2024-0024)
Unlike conventional image processing algorithms, μPIX adapts its training to the characteristics of the microscope, making it personalized and highly precise.
It improves image quality while preserving fine structures and intensity relationships, which is essential for quantitative imaging.
Thanks to its capacities, it extends the usefulness of old equipment, offering a cost-effective and sustainable alternative to replacement.
Better results than existing tools
In their publication, the authors show that μPIX outperforms both traditional denoising methods and popular deep learning tools such as CARE or Cellpose3.
It also improves downstream applications: using μPIX as a pre-processing step enhances segmentation accuracy by up to 3% compared to existing pipelines.
Reviving aging detectors
The team went one step further and applied μPIX to an ambitious task: restoring images from an outdated Multi-Alkali photodetector so they resemble those acquired with a high-performance GaAsP detector.
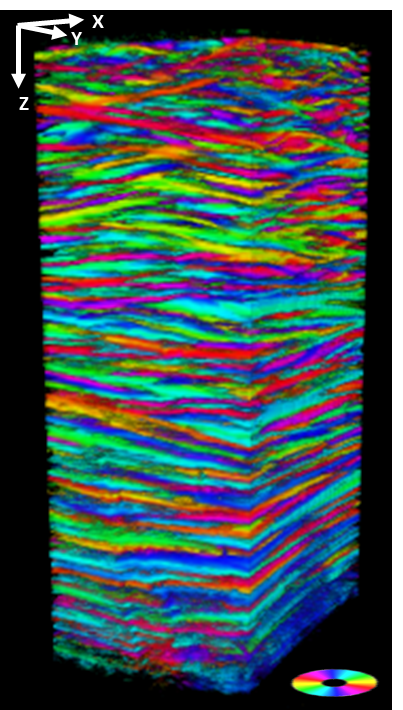
The results are impressive: μPIX manages to compensate for signal loss along the z-axis (represents the depth), recover structural information, and maintain a near-linear relationship between the predicted and original intensities, enabling quantitative analysis on images that would otherwise be considered obsolete.
From user-centered to hardware-centered AI
Unlike most AI tools that require users to train their own models, μPIX proposes a platform-centered paradigm: platforms train one model, tailored to their equipment, and provide it to their users. This approach reduces redundancy, improves consistency, and aligns with the principles of frugal and shared AI development.
The code and models are freely available on GitLab, and μPIX is already proving to be a useful asset for microscopy platforms seeking long-term performance with limited hardware budgets.
Read the full scientific article here: https://www.degruyterbrill.com/document/doi/10.1515/mim-2024-0024/html