A tight collaboration of research/development teams and microscopy research platforms
The Montpellier node comprises a tight collaboration of research/development teams and microscopy research platforms. The main objective of this FBI node is to develop, implement and provide access of state-of-the-art optical microscopy systems allowing microscopic imaging from unicellular organisms to whole animals. Specifically, our emphasis is on super-resolution and fluctuation microscopies, high-throughput high-content microscopies, and in vivo cellular imaging and manipulation. Our national specificity resides in the use of high-end microscopies to study different aspects of chromatin biology, such as chromosome organization and gene expression.
FBI-Montpellier is composed of a microscopy research and development facility (MARS of the CBS), two microscopy facilities (MRI and IPAM), and two R&D groups (Nollmann and Margeat labs). Several active and synergistic collaborations exist between these different entities in which R&D groups contribute their expertise in optical/instrument development and platforms contribute their savoir-faire in user service and in general logistics.
2021 in numbers
- 921 hosted projects
- 187 publications
- 6 training programs
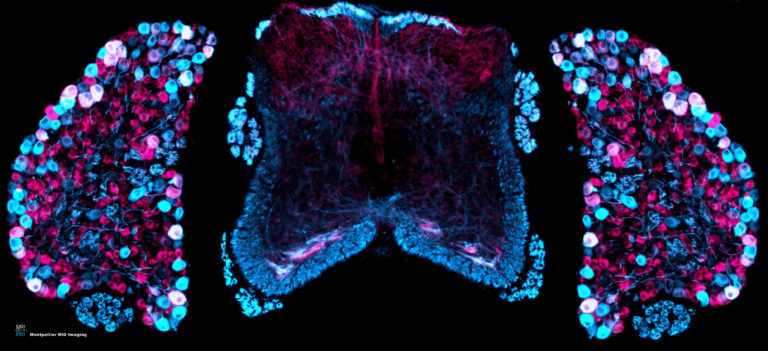
State-of-the-art Imaging
- Six PALM/STORM setups (3D, multicolor, multiplane, correlative)
- 3D-structural Illumination Microscope (OMX) fitted with PALM/STORM
- Five Fluorescence Correlation Spectroscopy and two Singel-Plane Illumination Microscopes
- High-content Screening and Tomographic 3D electron Microscopies
- Multiphoton microscopes for in vivo Intravital and Serial Block Face Imaging
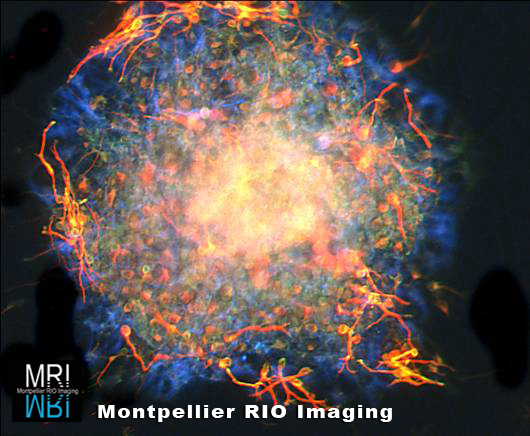
Technological innovations
- Number & Brightness (PNAS, 2012)
- smFISH (Nat. Methods, 2013)
- PALM/μfluidics (PLoS Biol., 2013)
- PIE-FCS (Nat. Comm, 2014)
- Multi-Focus Microscopy (patent filed 2015)
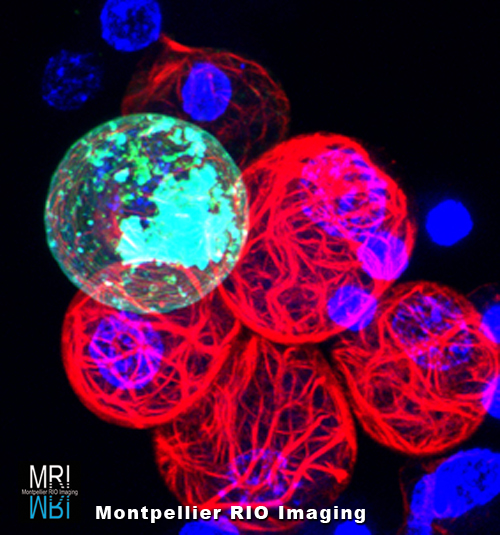
Training & Tech Transfer
- in 2021, 691 researchers received training in our facilities.
- France-BioImaging promoted the creation of the MARS facility, specifically devoted to tech transfer from R&D to platforms. https://fbimars.weebly.com/
Montpellier Resource Imaging

Facility: Montpellier Resource Imaging
Scientific director: Patrick Lemaire
Montpellier Ressources Imagerie (MRI) is a distributed imaging facility present on six sites in Montpellier (www.mri.cnrs.fr). MRI is labeled IBiSA and certified ISO9001-2008 LQRA. It has a staff of 30 engineers and is directed by P. Lemaire (CNRS). MRI manages numerous microscopes (36 photonic and 2 electron microscopes) and 14 analysis workstations, and especially microscopes for long term or short live experiments. MRI offers a complete set of state-of-the-art technologies, from single molecule to small organism imaging. The platform offers and develops 3D-SIM, SPIM, FCS/FCCS, CLEM and 2photons microscopies, and also develops a new service of High Content Screening, with a specific emphasis on gene expression analysis by smFISH techniques. MRI organizes regular training sessions with theoretical presentations and practical sessions about advanced light microscopy and image analysis. Once trained, a user can freely access microscopes on a pay-per-use basis. For the screening facility, the access is evaluated on a project-by-project basis.
Microscopy systems available @MRI
IPAM

Facility: IPAM
Head: Chrystel Lafont and Pierre Fontanaud
IPAM is a platform for the investigation of small animals. IPAM platform is under ISO9001 certification (starting from June 2014) and a labeled IBiSA facility. IPAM is headed by P. Mollard (CNRS, IGF) and with help of C. Lafont (tech leader). IPAM-IGF is dedicated to cellular in vivo imaging techniques in both anesthetized and vigile animal models. Our latest development involves 2-photon cellular in vivo microscopy with long-range objectives (Mitutoyo, wd: 2cm, x20 magnification, NIR transmission) readily applicable to imaging of deep tissues structures (metabolic brain, pancreatic islets from animal models of diabetes) in anesthetized animal models. Access to IPAM-IGF equipment is based on project selection (http://www.ipam.cnrs.fr/), IPAM-IGF is also an international member of the National Biophotonics and Imaging Platform Ireland (NBIPI, http://www.nbipireland.ie/ ).
Microscopy systems available @IPAM
PIBBS – MARS & AFM Facilities @CBS

Facility: PIBBS – MARS & AFM Facilities @CBS
Scientific directors: Christine Doucet (MARS) & Luca Costa (AFM)
PIBBS-MARS & AFM comprises two facilities :
- The AFM facility is the only one in the FBI Infrastructure to provide access to state-of-the-art Atomic Force Microscopes, including a custom built high-speed AFM.
- The MARS facility is devoted to cutting-edge optical microscopies such as Single Molecule Localization Microscopy, smFRET, PIE-FCCS, 2-photons FCS, single particle tracking, etc… The MARS R&D division, closely associated to two R&D teams, is in charge of implementing and developing new custom
advanced microscopies (such as STED-FCCS, 2foci-FCS or multifocal microscopy) before their transfer to the facility.
Moreover, the AFM and MARS facilities and staff work closely together to develop and transfer new correlative imaging modalities, such as AFM / Superresolution or AFM / Confocal (FLIM, spFRET).
Users are assisted by dedicated research engineers and scientific coordinators to define the best approach, experimental design, help in data acquisition and analysis.
http://www.cbs.cnrs.fr/index.php/fr/fluorescence
Microscopy systems available @PIBBS
Mechanisms of DNA segregation and remodelling Team @CBS

R&D team: Mechanisms of DNA segregation and remodelling Team @CBS
Head: Group Leader: Marcelo Nollmann
We develop single-molecule and advanced microscopy methodologies to investigate the mechanisms underlying DNA segregation and remodeling in live cells.
Our current research projects:
- DNA organization and segregation in bacteria
- Eukaryotic DNA structure
- Molecular Motors
- Single-molecule & advanced optical microscopies
Integrated Biophysics of Membranes Team @CBS

R&D team: Integrated Biophysics of Membranes Team @CBS
Group Leaders: Emmanuel Margeat & Pierre-Emmanuel Milhiet
Our research aims at characterizing macromolecular complexes governing major biological processes, focusing on transcription regulation, signaling and remodeling of biological membranes. To achieve these goals, we develop, combine and use advanced single molecule biophysical methods (such as atomic force and fluorescence microscopies), as well as DNA nanotechnology.
Research:
Structure and dynamics of nucleoproteic and membrane assemblies
ICAR Research-Team @LIRMM

R&D team: ICAR Research-Team @LIRMM
FBI Contact: Emmanuel Faure
The ICAR Research-Team develops research themes combining the interaction and processing of visual data such as 2D, 3D, videos or nD+t image sequences and 3D meshes. The team is structured according to 3 research axes: Analysis & Processing which is focused on new low-level processing techniques of visual information, Multimedia Security which is interested in securing visual data and Modeling & Visualization which aims at representing large and complex visual data sets.
Expertise of the Team
- The Analysis & Processing axis is interested in new low-level information processing techniques to improve the information perceptible in the image and to take into account, within the same theoretical framework, the imprecise, uncertain and incomplete (the different types of error in
visual data processing). - The Multimedia Security axis is interested in the security of visual data. In order to ensure this security, coding algorithms are developed combining tattooing, steganography, forensics, encryption and authentication and often requiring robustness to compression.
- The objective of the Modeling & Visualization axis is to model large sets of complex data (in dimension and nature) in order to allow intuitive visualization or to manipulate these data to extract knowledge from them.
RNA biogenesis @IGH

R&D team: RNA biogenesis @IGH
Team Leader: Edouard Bertrand
Our group has strong interests in gene expression mechanisms, from transcription to translation. While we are interested in the regulation of these processes and their functional consequences, the big question that moves us is to understand how they occur in the context of a living cell.
Indeed, cells are not only the individual units where gene regulation takes place, but they are also incredible objects: if we consider RNA and proteins, a typical cell contains several hundreds of thousands of different molecular species, with some present in millions of copies per cell while others in only few. In order to function with such a high complexity within a crowded molecular environment, cells rely on two main tools: (i) chaperones specialized in the control of molecular interactions; (ii) a remarkable degree of spatial organization, which also allows a high plasticity and a high dynamics of molecules. It is to get insights into these very fundamental questions that we first developed tools to image single mRNAs in live cells. With these tools in hands, and others that we developed later, we aim at imaging the basic mechanisms of gene expression directly in living cells, thereby providing a renewed vision of these fundamental processes.
Our strategy is to invest in methodological developments to access and image new facets of gene expression, usually at the levels of single molecules. These developments are mostly focused on imaging RNA metabolism and they are guided by our current scientific questions.
Methodological developments require multidisciplinary approaches, and we have therefore developed a stable network of collaborators who complement our own expertise. This includes the groups of: (i) C. Zimmer and F. Müller (Pasteur Institute, Paris; https://research.pasteur.fr/en/team/imaging-and-modeling/), a physicist team with a great expertise in image analysis; (ii) T. Walter (Curie/Ecole des Mines; Paris; http://members.cbio.mines-paristech.fr/~twalter/), an applied mathematician expert in high-content microscopy and in complex, high-dimensional dataset analysis; (iii) O. Radulescu (Montpellier University; https://systems-biology-lphi.cnrs.fr/), a mathematician expert in modeling biological processes. More recently, we initiated collaborations with chemists to develop novel RNA probes and biosensors.
Our group works in three main areas: transcriptional and translational regulation, as well as chaperone-mediated control of molecular interactions.
Recent publications
The mechanism of force transmission at bacterial focal adhesion complexes. Nature, 2016 Nov 24;539(7630):530-535. doi: 10.1038/nature20121.
Plasmonic Nanoantennas Enable Forbidden Förster Dipole-Dipole Energy Transfer and Enhance the FRET Efficiency. Nano Lett. 2016 Oct 12;16(10):6222-6230.
A single-molecule view of transcription reveals convoys of RNA polymerases and multi-scale bursting. Nat Commun. 2016 Jul 27.
Highly efficient multicolor multifocus microscopy by optimal design of diffraction binary gratings Scientific Reports, in press.
Bacterial partition complexes segregate within the volume of the nucleoid Nature Communications, 7: 12107, 5 July 2016
Condensin- and Replication-Mediated Bacterial Chromosome Folding and Origin Condensation Revealed by Hi-C and Super-resolution Imaging. Molecular Cell 59 (4): 588-602, 20 August 2015
Fine tuning of sub-millisecond conformational dynamics controls metabotropic glutamate receptors agonist efficacy. Nature communications”>Nat Commun. 2014 Oct 17;5:5206.
FISH-quant: automatic counting of transcripts in 3D FISH images. Nat Methods. 2013 Apr;10(4):277-8.