Atomic Force Microscopy (AFM) is a scanning probe microscopy technique that relies on measuring the interaction forces between a sharp tip and the surface of a sample to generate high-resolution images of its surface features and mechanical properties. A very broad range of sample types can be imaged with this technique at a very high resolution – at sub-nanometer level for some of them! Discover the AFM at the Montpellier node of France-BioImaging with Christine Doucet from Integrative Biophysics of Membranes team of the Centre de Biochimie Structurale.
Quickly visualize dynamic biological processes with High-Speed AFM
AFM provides images in physiological conditions, in liquid, over a length-scale ranging from few nanometers (single biomolecules) to tens of micrometers (living cells). In fact, the resolution depends on the tip radius and sample properties. For some of them, you can routinely obtain a nanometer lateral resolution and Angstrom axial resolution!
You want a video-rate version of the biological samples you are imaging? The High-Speed AFM, permits the acquisition of movies at approximately 10 images per second, enabling the visualization at nanoscale of dynamic biological processes involving biomolecular interactions, diffusion or conformational changes. It delivers nanometric resolved images typically at the same speed as conventional fluorescence microscopes!
Unravel the chemical information of your sample by combining AFM with…
AFM in ambient conditions and in liquids has a key limitation in that it does not directly provide chemical information about the sample being imaged. However, this limitation can be overcome by combining AFM with other techniques to obtain additional information about the sample’s composition.
One commonly used technique in correlation with AFM is fluorescence microscopy. This combined approach of fluorescence labeling and AFM provides valuable insights into the chemical and biological properties of the sample. It was recently used on the Montpellier custom-made correlative AFM / fluorescence setup to observe the sublocalization of proteins in HIV-1 budding sites 1. They also used it to unambiguously attribute some unexpected configurations of the nucleoplasmic sides of Nuclear Pore Complexes 2. In these two cases, fluorescently-labeled proteins were imaged by dSTORM (direct STochastic Optical Reconstruction Microscopy). Of note, the lateral resolution of dSTORM and AFM are both in the 20 nm range with such samples, which makes their combination ideal!
In addition to fluorescence microscopy, AFM can also be correlated with other complementary techniques to obtain chemical information about the sample, such as Raman spectroscopy, Infrared Spectroscopy, X-Ray spectroscopy, microscopy and scattering.
Learn more about AFM applications
Here are 2 studies where Atomic Force Microscopy were essential:
- Structure and mechanics of the human nuclear pore complex basket using correlative AFM-fluorescence superresolution microscopy
Combining mechanical and superresolution measurements to reveal the plasticity of the Nuclear Pore Complexes
Nuclear pore complexes (NPCs) are the only gateways between the nucleus and cytoplasm in eukaryotic cells, facilitating the transport of selected cargoes of size from a few up to hundred nanometers. This versatility implies an important pore plasticity. Here, by combining atomic force microscopy (AFM) and single molecule localization microscopy (SMLM), a group led by France-BioImaging R&D team members Christine Doucet and Pierre Emmanuel Milhiet revealed that the NPC basket is very soft and explores a large conformational landscape: apart from its canonical basket shape, it dives into the central pore channel or opens, highlighting how this structure can adapt, and let morphologically diverse cargoes shuttle through NPCs.
Vial et al., Nanoscale, 15, 5756-5770 (2023)
- The structure of pathogenic huntingtin exon 1 defines the bases of its aggregation propensity
Structural Biology meets Correlative Imaging
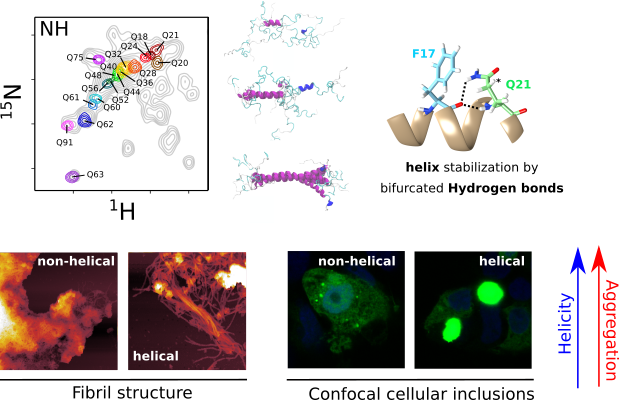
Huntington’s disease is a neurodegenerative disorder caused by an extended polyglutamine (poly-Q) tract in huntingtin. Here, using NMR, the team of Pau Bernado (CBS Montpellier) demonstrated that this poly-Q tract adopts long α-helical conformations. By adding correlative Atomic Force Microscopy and Fluorescence Microscopy data obtained in the FranceBioImaging facility PIBBS in Montpellier, they could demonstrate that the stability of this α-helix is a stronger signature than the number of glutamines, in defining the aggregation kinetics and the structure of the resulting fibrils, potentially linked to their pathogenicity.
Elena Real et al., Nature Structural & Molecular Biology, 30, 309–320 (2023)
How to use Atomic Force Microscopy at France-BioImaging?
Atomic Force Microscopy is open to collaborations under Proof-of-concept studies via Euro-BioImaging webportal (www.eurobioimaging.eu/service)! At the Montpellier node of France-BioImaging, you will be in contact with Dr Luca Costa (costa@cbs.cnrs.fr) with whom you will talk about the feasibility and the inherent experimental constraints linked to the technique. The collaboration procedure is discussed on a case-by-case basis, depending on the duration and technicity of the required experiments. Feel free to submit your project!
Thanks to Christine Doucet and Emmanuel Margeat for providing helpful information!
1. Dahmane, S., Doucet, C., Le Gall, A., Chamontin, C., Dosset, P., Murcy, F., Fernandez, L., Salas, D., Rubinstein, E., Mougel, M., et al. (2019). Nanoscale organization of tetraspanins during HIV-1 budding by correlative dSTORM/AFM. Nanoscale 11, 6036–6044.
2. Vial, A., Costa, L., Dosset, P., Rosso, P., Boutières, G., Faklaris, O., Haschke, H., Milhiet, P.-E., and Doucet, C.M. (2023). Structure and mechanics of the human nuclear pore complex basket using correlative AFM-fluorescence superresolution microscopy. Nanoscale 15, 5756–5770.